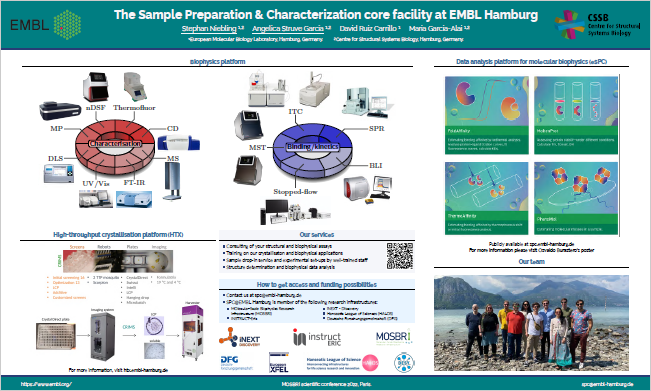
6) EMBL-SPC – Sample Preparation and Characterisation Facility, EMBL Hamburg (HH)
Location: EMBL, DESY campus, Hamburg, Germany
Web-site(s):
MOSBRI reference site for the following techniques:
DSF: Differential Scanning Fluorimetry, BLI: BioLayer Interferometry
Practical information:
View the practical information for EMBL-SPC here

Partner description
The EMBL-SPC facility is located next to an integrated infrastructure for structural biology at the PETRA-III ring at DESY Hamburg, open to the international research community. EMBL operates three beamlines for applications in X-ray crystallography and small angle X-ray scattering. The biophysical platform of the EMBL-SPC includes cutting-edge technologies to measure interactions and to precisely determine the stability, shape and size of different biomolecules and biomolecular assemblies. The characterisation of these structures are studied by studying parameters such as stability, shape, folding state, aggregation tendency, homogeneity and oligomeric states. The interactions in which they are involved can be followed by measuring biophysical criteria like; specificity, kinetics, affinity, and thermodynamics.
Instruments / methodologies offered through TNA
Biophysical characterization of Integral Membrane Proteins by High-throughput stability screening: Protein stability in detergent or membrane-like environments is the bottleneck for structural studies on integral membrane proteins (IMP). Our pipeline allows high-throughput screening to identify optimal conditions that are suitable for downstream handling during purification. It includes thermal-denaturation assays monitored by differential scanning fluorimetry, static and dynamic light-scattering, and infrared spectroscopy (attenuated total reflection). We have developed screens to identify optimal detergents, buffers and additives for solubilisation. Lipids play a major role in preserving the native environment of eukaryotic membrane proteins and our pipeline helps to detect which lipids or cocktail-lipids are necessary to keep the integrity of IMPs. We address the functionality of membrane transporters by using a setup of solid-supported membrane (SSM) electrophysiology based on the SURFE²R technology for measuring direct transport of substrates across the lipid bilayer.
Interaction studies: We offer a compilation of methods to quantify protein-ligand interaction. This includes isothermal calorimetry, biolayer interferometry (Octet), surface plasmon resonance (Biacore) and differential scanning fluorimetry (nDSF). For the latter technique, we offer an analysis webserver that uses isothermal analysis in order to obtain binding affinities.
Time-resolved studies: This pipeline offers several methods to obtain kinetic information from different read-outs:
- time-resolved SAXS beamline (P12).
- stopped-flow device that allows read out of absorbance, fluorescence and fluorescence anisotropy.
- binding kinetics of surface-immobilized proteins based on biolayer interferometry (Octet).
We offer access to
- Determination of purity,
- protein identification and intact mass determination,
- dispersity analysis,
- screening of conditions by thermal denaturation assays,
- protein secondary structure determination and temperature stability by infrared and circular dichroism spectroscopy,
- FT-IR spectrometry of coated nanoparticles,
- molecular modelling, and
- MD simulations and analysis of trajectories.
Access modality: Access is typically allocated as 4 days. Both physical access and remote access are offered.
Sample requirements and further technical details
Full details of the sample requirements for each of the techniques offered at this TNA site can be found here. Further, more specific details, of some of the techniques offered are also included.

